For most research projects, it is important to anonymize data from human subjects: no names, birth dates, addresses in file names or metadata. But this is not enough because the anatomical data, reconstructed in surface with the skin allows to identify the subject. It is therefore necessary to “deface” these anatomical images, which means to blur or remove the voxels of the face, while preserving the voxels of the brain which will be used for data analysis. This defacing is actually imposed by some data sharing platforms such as OpenNeuro.
There are several software solutions to do this defacing automatically. Here are some illustrations in pictures:
| Original | pydeface | mri_deface | quickshear | deedeface |
|---|---|---|---|---|
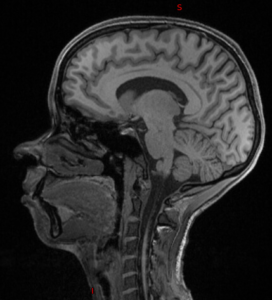 | 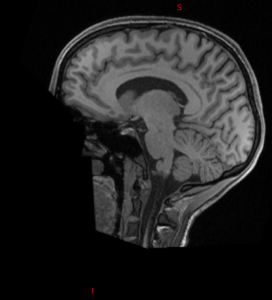 | 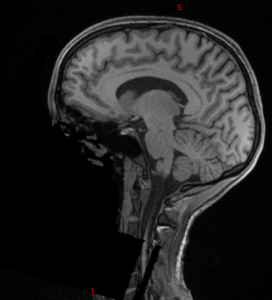 | 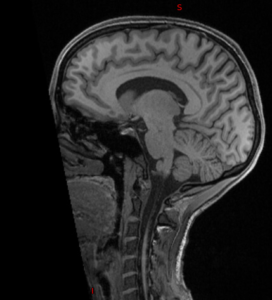 | 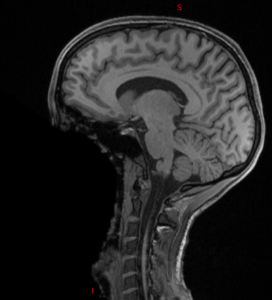 |
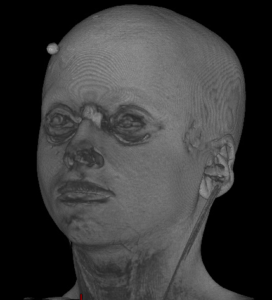 | 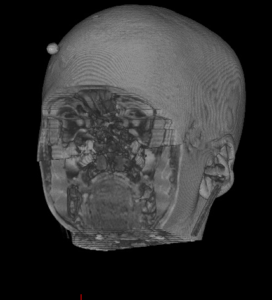 | 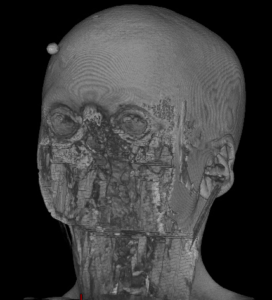 | 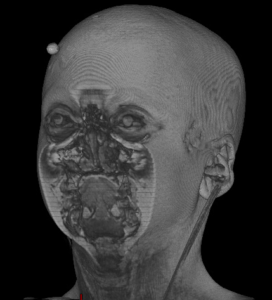 | 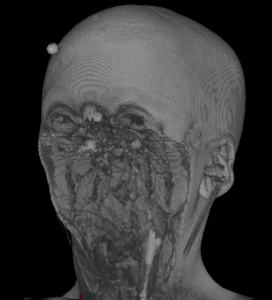 |
An example implementation is possible thanks to BIDSApp Bidsonym.
The MRI center has tested the different defacing software available in this BIDSApp, namely:
– pydeface ( https://github.com/poldracklab/pydeface )
– mri_deface ( https://surfer.nmr.mgh.harvard.edu/fswiki/mri_deface )
– quickshear ( https://www.usenix.org/legacy/event/healthsec11/tech/slides/schimke.pdf )
– deepdefacer ( https://github.com/AKhazane/DeepDeface )
– mridefacer ( https://github.com/mih/mridefacer )
These tests were conducted using a bidsonym singularity image installed on the mesocenter.
Images acquired from one research project (T1w @ 0.8mm iso acquired on the MRI Center’s 3T MRI) were used for this test (N=40).
Example of observed measurements (calculated by MRIQC):
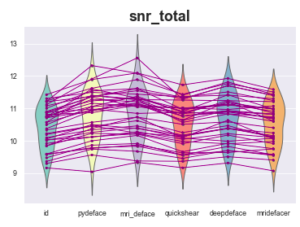
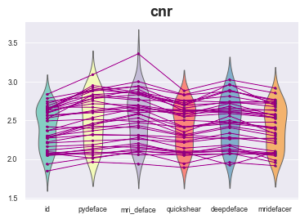
At first glance, everything is fine, the measurements seem relatively homogeneous between the original image and the defaced images provided by the different defacing algorithms.
However, one measurement looks fishy:
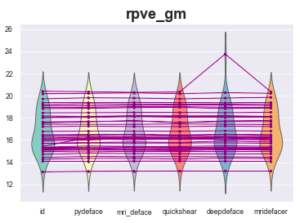
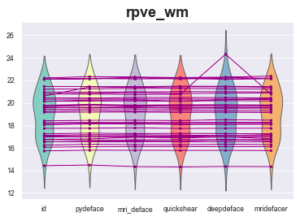
Definition: RPVE: Computes the residual partial voluming error of each tissue class: Lower values are better.
Who is this intruder? When looking at which subjects deviates from the original image for this measurement, we spot sub-19
Let’s have a look at the image for this subject sub-19:
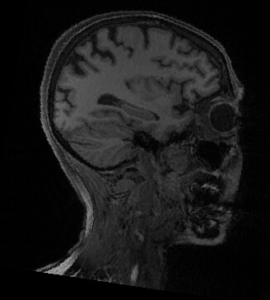 | 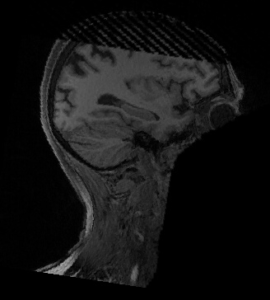 |
Indeed, there was a problem ! Deepdefacer is fast but not always robust !
Morality: always check the data at each stage of processing …


